Subscribe to Pittwire Today
Get the most interesting and important stories from the University of Pittsburgh.Last summer, undergraduate research assistant Gabrielle Gentile watched her colleagues in the University of Pittsburgh Hatfull Lab work diligently to identify viruses that could infect and kill the deadly bacteria Mycobacterium abscessus.
Gentile (A&S ’18) knew the hunt for the right bacteria-killing virus, also known as a phage, involved work she and other students were doing in SEA-PHAGES, a national program that teaches undergraduate science students the process of discovering phages in soil and other natural environments. She knew the Hatfull Lab was asking student phage hunters to submit their discoveries for review.
What she didn’t know was the killer phages they were seeking would save the life of a 15-year-old with cystic fibrosis whose recovery from a lung transplant had been complicated by an antibiotic-resistant infection.
“It was really exciting to hear about that. I know among my entire SEA-PHAGES cohort there was excitement on social media,” said Gentile, who is currently enrolled in the University of North Carolina School of Medicine’s Biomedical and Biological Sciences PhD program.
“This happened with our lab — it’s not every day your lab is involved in something that saved a life.”
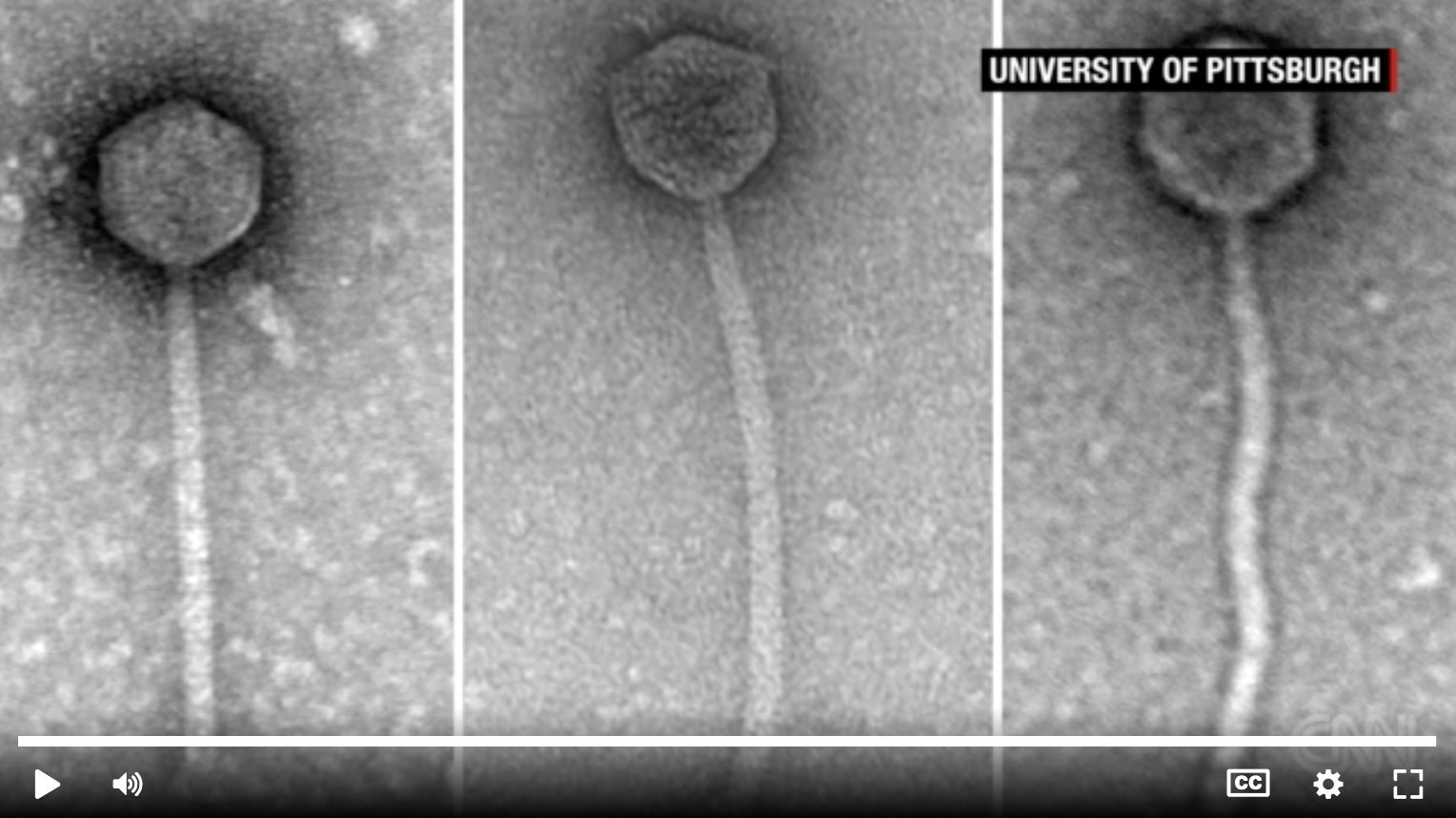
Graham Hatfull, the Eberly Family Professor of Biotechnology and SEA-PHAGES lead scientist, led the team that identified and engineered the phages to create the phages cocktail. The cocktail was designed to treat a 15-year-old cystic fibrosis (CF) patient named Isabelle Carnell-Holdaway who had a double lung transplant and was suffering from a life-threatening infection with an antibiotic resistant strain of M. abscessus. The cocktail was administered intravenously twice daily and topically to her surgical wound and to skin lesions caused by the infection.
The lifesaving research from Hatfull’s lab is the first known attempt to use phages to treat a mycobacterial infection in a human being, as well as the first therapeutic use of engineered phages.
“This is a very exciting development,” said Hatfull. “These types of infections are common in CF patients and are extremely difficult to treat due to antibiotic resistance. Although phages may not yet be broadly applicable for such infections, this study shows that it may be a highly effective treatment.”
Hatfull and Helen Spencer, a respiratory pediatrician and clinical lead for lung transplant services at London’s Great Ormond Street Hospital, outlined the process in a paper published in Nature Medicine on May 8.
The full team includes Hatfull Lab research associates Rebekah Dedrick and Carlos A. Guerrero-Bustamante, research assistant Rebecca Garlena, visiting lab instructors Daniel Russell and Deborah Jacobs-Sera, Robert “Chip” Schooley from the University of California San Diego Department of Medicine, and Katrina Ford, Kathryn Harris, Kimberly Gilmour and James Soothill from Great Ormond Street Hospital.
Behind the research
The national contingent of students that make up SEA-PHAGES, which stands for Science Education Alliance-Phage Hunters Advancing Genomics and Evolutionary Science, played a critical role. In order to find the phages that made the lifesaving cocktail, the team examined a collection of approximately 10,000 mycobacteriophages, 1,700 of which had undergone genetic sequencing, and isolated three phages capable of infecting Isabelle’s antibacterial resistant M. abscessus strains.
That collection of mycobacteriophages came directly from the efforts of approximately 20,000 undergraduate SEA-PHAGES participants at more than 130 institutions who have discovered more than 15,000 over the last decade. The program is administered by Hatfull and his team and the Howard Hughes Medical Institute’s Science Education Division. At Pitt, 362 students helped in the hunt this academic year.
Through SEA-PHAGES, students are tasked with finding the viruses hidden within soil, identifying them in the lab and giving them names before they are entered into the larger collection. The phages used to create Isabelle’s cocktail were named Muddy, ZoeJ and BPs.
“My very first phage was Vincenzo,” said Gentile. “I had many subsequent ones with all types of names.”
Beyond discovery, students learn advanced scientific techniques that include DNA sequencing and analysis of gene function. The experience was enough to sway Gentile, who had been on a path toward becoming a physician before the program.
“Before Pitt I was interested in biology, but SEA-PHAGES steered me toward the research and discovery route. It showed me the importance of asking scientific questions and discovering the answers for yourself — and having the satisfaction of making that discovery,” she said.
Gentile said Vincenzo and her other phages weren’t part of the groundbreaking trial, but that being part of the team “brought tears of joy.”
“It was very heartening reading the articles and knowing the patient has resumed routine activities and is living a life all because of phages a student discovered.”
Hatfull Lab goes viral
News of the Hatfull lab’s lifesaving international collaboration spread fast. Read some of the highlights:
A Dying Teenager’s Recovery Started in the Dirt
The Atlantic
Genetically Engineered Viruses Treat Antibiotic-Resistant Infection
The Wall Street Journal
How Genetically Engineered Viruses — and a Rotten Eggplant — Prolonged a Teenager's Life
STAT News
Phage Viruses Tailored in Pitt Lab Used to Treat Patient With Drug-resistant Infection
Pittsburgh Post-Gazette
Teenager Recovers From Near Death in World-first GM Virus Treatment
The Guardian
Genetically Tweaked Viruses Just Saved a Very Sick Teen
Wired
Viruses Genetically Engineered to Kill Bacteria Rescue Girl with Antibiotic-resistant Infection
Science magazine
An Antibiotic Alternative? Using a Virus to Fight Bacteria
U.S. News & World Report
Genetically Modified Viruses Help Save A Patient With A 'Superbug' Infection
NPR