| front |1 |2 |3 |4 |5 |6 |7 |8 |9 |10 |11 |12 |13 |14 |15 |16 |17 |18 |19 |20 |21 |22 |23 |24 |25 |26 |27 |28 |29 |30 |31|32 |33 |34 |35 |36 |37 |38 |39 |40 | 41 |42 |43 |review |
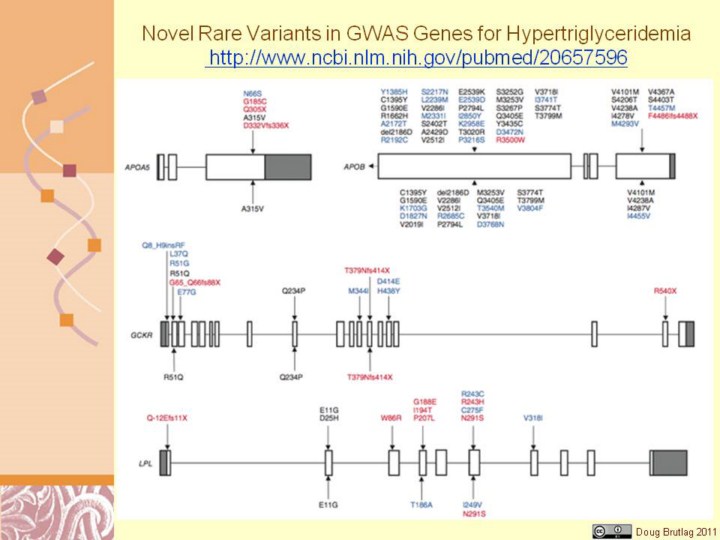 |
http://www.ncbi.nlm.nih.gov/pubmed/20657596 Figure 1 Rare variants identified by resequencing GWAS-identified genes in individuals with HTG and controls. Variants listed above gene maps were identified in affected individuals, and those below gene maps were identified in controls. Rare variants are colored as follows: black, identified in control subjects or previously identified in subjects of unknown clinical status; blue, exclusive to affected individuals or controls; red, proven biological dysfunction or truncation. Nomenclature for variants refers to functional protein sequences. Only exons 26 and 29 were resequenced in APOB. Gene maps are roughly to scale, although they differ in scale between genes. |