| front |1 |2 |3 |4 |5 |6 |7 |8 |9 |10 |11 |12 |13 |14 |15 |16 |17 |18 |19 |20 |21 |22 |23 |24 |25 |26 |27 |28 |29 |30 |31 |32 |33 |34 |35 |36 |37 |38 |39 |40 |41 |42 |43 |review |
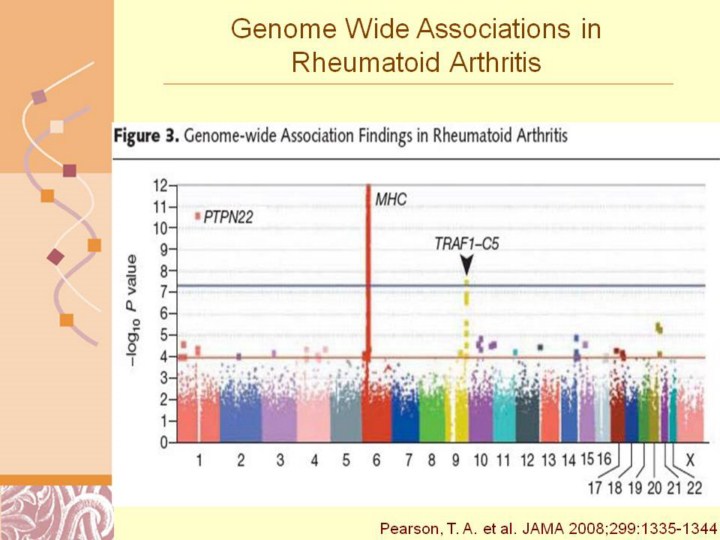 |
Genome-wide association studies assume a priori hypotheses about candidate genes or regions that might be associated with disease; rather, they test single-nucleotide polymorphisms (SNPs) throughout the genome for possible evidence of genetic susceptibility. Associations plotted as −log10 P values for a genome-wide association study in 1522 cases with rheumatoid arthritis and 1850 controls, showing single data points for SNPs with P 10−4 (lower horizontal red line) for 22 autosomes and the X chromosome. The predefined level of significance, at 510−8 is shown with a horizontal blue line. SNPs at PTPN22 on chromosome 1, the major histocompatibility comples (MHC) on chromosome 6, and the TRAF1-C5 locus on chromosome 9 exceed this threshold. Reproduced with permission from Plenge et al.47 |